OnkoGenetiks™ analyzes 486 genes associated with cancer with Molecular Profiling. OnkoGenetiks Molecular Profiling test determines the genetic changes that cause cancer progression and provides a personalized treatment solution with a detailed report on all possible treatment alternatives. This test also carries the feature of being a potential biomarker for cancer therapy by analyzing the Tumor Mutation Burden and the total number of mutations in the tumor genome.
What is Molecular Profiling?
Detection of cancer-related biomarkers has become the fundamental step for an individualized treatment approach in cancer patients. "OnkoGenetiks ™ Molecular Profiling" panel, designed for this purpose, offers a comprehensive genomic profiling analysis of formalin-fixed and paraffin-embedded (FFPE) or fresh tumor tissue samples. “OnkoGenetiks ™ Molecular Profiling” is used to identify DNA variants in different tumor types, as well as to detect Tumor Mutation Burden (TMB) and Microsatellite Instability (MSI), which are immunotherapy biomarkers.
By making use of next-generation sequencing (NGS) technology, this comprehensive test panel, which allows simultaneous screening of variants in 486 genes, can provide fast and comprehensive results even in cases where the sample amount is low.
As a result of the data obtained from OnkoGenetiks™ Molecular Profiling and TMB panel, it is possible to evaluate the following options:
- Targeted therapy
- Personalized treatment
- Smart drug / immunotherapy
Why OnkoGenetiks™ Molecular Profiling?
- Determination of germline mutations through Molecular Profiling performed from blood, thus enabling a real TMB calculation
- Detection of DNA variants in 486 genes, Microsatellite Instability (MSI) and Tumor Mutation Burden (TMB),
- Determination of personalized treatment options,
- Possibility of avoiding the potential side effects of chemotherapy,
- Availability of results in a short time (15-21 working days),
- High sensitivity and read depth (utilization of Unique Molecular Index-UMI),
- Access to more than 10 million biomedical findings and over 1 million variant-phenotype relationships by making use of QIAGEN bioinformatics solutions. (All data belonging to the patients is stored and analyzed at the Genetiks laboratory in Turkey.)
What is Tumor Mutation Burden (TMB)?
What is the principle of TMB analysis?
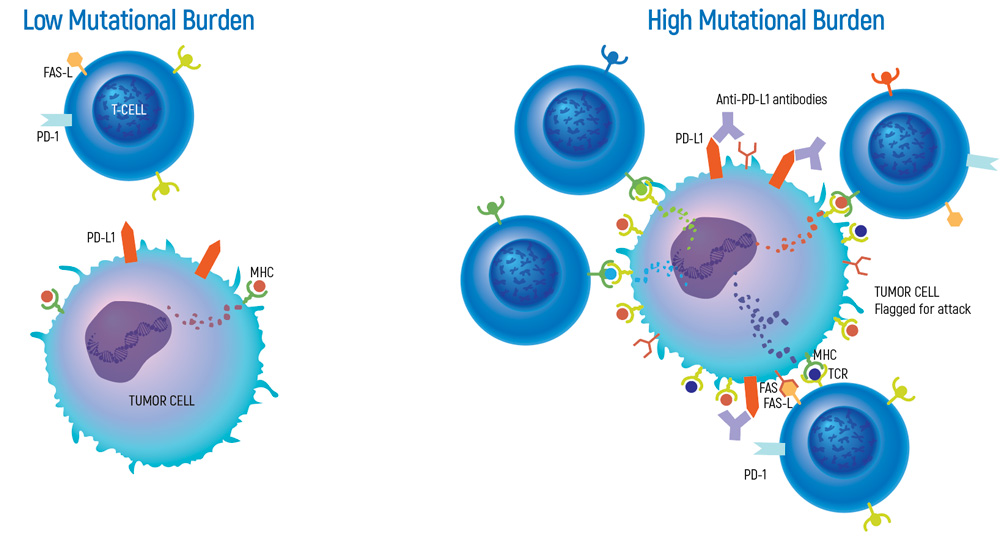
Tumors with high mutation burden have the potential to generate a larger number of neoantigens, making them more immunogenic. Neoantigens are different than the proteins normally synthesized by our body, therefore they are perceived as extraneous and lead to an antitumor response by the immune system.
Cancer immunotherapies have the potential to treat cancer by taking advantage of the power of the immune system. However, only 20-40% of cancer patients respond to treatments based on immunotherapy. TMB can help to identify patients who are more likely to benefit from cancer immunotherapies for certain tumor types. Identifying these patients prior to immunotherapy is important in terms of time that is critical in the treatment process, cost of treatment and potential side effects.
The relationships between TMB levels and the response to immunotherapy treatments have been demonstrated in various studies.
- In a study of anti-PD-1 immunotherapy in lung cancer, high TMB was associated with 4 times longer progression-free survival (PFS), compared to low TMB.
(Rizvi et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. 2015. Science. 348 (6230):124-128.) - In melanoma patients receiving anti-CTLA-4 immunotherapy, high TMB was associated with 3.5 years longer overall survival (OS), compared to low TMB.
(Snyder et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. 2014. NEJ Medi. 371:2189-2199.) - A significant correlation has been indicated between TMB and progression-free survival and objective response rates for NSCLC, melanoma, esophagogastric cancer, SCCHN, and RCC.
(Samstein et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. 2019. Nature Genetics, 51, 202–206.)

What is MSI (Microsatellite Instability)?
Microsatellites are DNA repeat sequences that appear to be in a stable state throughout the life of an individual and whose functions in the genome are not fully revealed. Microsatellite Instability (MSI), is the condition of accumulating variations in the tumor DNA due to deletion or insertion in microsatellite repeats as a result of inactivation in MMR (mismatch repair) genes. In this case, microsatellites observed in the DNA of specific cells (such as cancer cells) differ from the DNA repeats in normal cells of the same individual.
DNA mismatch repair (MMR) system is one of the mechanisms the cell uses to repair DNA damage. In the case that there are mutations in the genes encoding MMR enzymes, mismatches in DNA sequences cannot be repaired, and erroneous DNA sequences called microsatellites build up, creating genomic instability. Microsatellite Instability (MSI) caused by such deficiencies in DNA mismatch repair, occurs frequently in certain tumor types.
How to determine MSI (Microsatellite Instability)?
If a tumor shows microsatellite instability (MSI), this can be detected by comparing microsatellites in the tissue sample of the tumor with a normal tissue from the same person. Genomic DNA is isolated from both tumor and EDTA-blood sample, and PCR analysis is performed using a panel containing five different microsatellite markers (BAT25, BAT26, D5S346, D2S123, D17S250) determined by the National Cancer Institute. The PCR products are then subjected to fragment analysis by capillary electrophoresis method and the results are evaluated in terms of microsatellite instability.
Tumors in which instability is observed for 2 or more of the 5 microsatellite markers analyzed (more than 30-40%) are defined as MSI-H (high-MSI). Tumors with instability in only 1 (less than 30-40%) of the 5 microsatellite markers analyzed are defined as MSI-L (low-MSI). Tumors with no instability observed in any of the microsatellite markers are also considered microsatellite stable (MSS).
Genetiks is accredited from CAP (College of American Pathologists) for MSI testing.


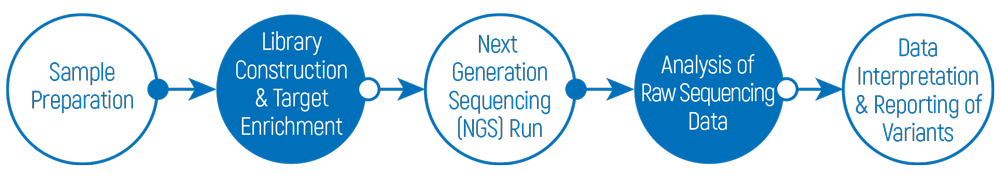
Overview of the general workflow for OnkoGenetiks TMB & MSI analysis
Microsatellite Instability (MSI) panel contains 7 markers corresponding to the MMR (Mismatch Repair) genes. MSI analysis is performed with ABI 3500xL Genetic Analyzer device and results are computed using the GeneMapper software.
Checkpoint inhibitor (CPI) therapy, which is a form of cancer immunotherapy, has demonstrated a remarkable clinical benefit in recent years for many cancer types such as skin, lung, bladder, kidney and in tumors having deficiency of MMR (Mismatch Repair).
Cancer immunotherapy emerged to be a very promising and powerful tool for the treatment of cancer. However, the ability to successfully identify patients who will benefit from this kind of therapy is still limited.
Tumor mutational burden (TMB), a measure of the number of somatic mutations per coding area of tumor genome, is a putative biomarker of response showing great promise in CPI and immunotherapy combination trials.
How is OnkoGenetiks™ Molecular Profiling & TMY Analysis Test done?



